P(1B)-Type Cu(I) Transporting ATPases ATP7A and ATP7B
From Proteopedia
Copper plays an important biological role in Homo sapiens, and is absolutely necessary as a co-factor for many proteins, including redox proteins, but can also participate in biologically harmful redox reactions if not regulated. As a result of the biological necessity of copper in Homo sapiens but inherent risks accompanying copper's presence in the body, its uptake, distribution and excretion is tightly regulated through specialized proteins. One such group of specialized proteins are the P(1B)-Type Cu(I) Transporting ATPases, which are named ATP7A and ATP7B [1]. The proteins ATP7A and ATP7B are specialized to bind Copper(I), and couple the energy released with the hydroplysis of the gamma phosphate of ATP to transport the Copper(I) across membranes [1]. In Homo sapiens these two proteins are essential for maintaining Copper(I) homeostasis within the body. Mutations in ATP7A, or ATP7B result in improper function of the proteins which has severe consequences on Copper(I) regulation within the body, disturbing the homeostasis. The diseases resultant of mutations in ATP7A or ATP7B are Menkes Disease, and Wilson's Disease respectively which are both inheritable, causing dysfunction of copper(I) homeostasis [1].
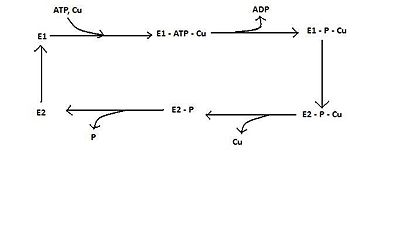
GeneticsThe protein ATP7A is coded for by the protein-coding gene sequence ATP7A in the human genome, which is located on the X chromosome [2]. The genomic sequence for ATP7A contains 23 exons with a total genomic length of approximately 140 kb [2]. The mature mRNA for ATP7A has a total length of approximately 8.5 kb, and is translated into an alpha polypeptide 1500 amino acids in length [2]. The protein ATP7B is coded for by the protein-coding gene sequence ATP7B in the human genome, which is located on chromosome 13 [3]. ATP7B's genomic sequence contains 21 exons and has an approximate total genomic length of 80 kb [3]. The mature mRNA for ATP7B has an approximate length of 6.5 kb, and its translated product is a beta polypeptide 1465 amino acids in length [3]. Structure and FuctionOverviewThe specialized Copper(I) transporting proteins ATP7A and ATP7B are a part of the family of P-Type ATPases. Even though ATP7A and ATP7B belong to a subfamily known as the P(1B)-Type ATPases, they share the same basic "core" architecture with the whole P-Type ATPase family [1]. For a reference of the "core" architecture of ATP7A and ATP7B, the complete structure of the Ca(2+) P-Type ATPase can be utilized (PDB code 3ar2). The "core" architecture of a P-Type ATPase consists of membrane spanning region, and a hydrophilic region protruding into the cytosol of the cell [1]. The first of the domains is the . The A-Domain is located near the Amino Terminal of the ATPase, and is the smallest of the cytoplasmic domains [4]. Next, is the . The P-Domain can be reversibly phosphorylated, and functions to become phosphorylated with the hydrolysis of a bound ATP [4]. The domain that contains the ATP-binding site of the P-Type ATPase is the . The N-Domain functions to bind ATP for its future hydrolysis to drive the transport of the heavy metal across the membrane [4]. The final domain is the . The M-Domain or transmembrane region consists of transmembraneous helices which form the intramembranous channel which provides a channel for the metal to be transported across the membrane [4]. In summary, the hydrophilic region of the protein protruding into the cytosol is made up of which form the intramembranous channel. The above is only the basic "core" architecture of all proteins in the family of P-Type ATPases, below the specific properties of each of these regions will by discussed for the sub-family P(1B)-Type ATPases ATP7A and ATP7B. In the following structural analysis of P(1B)-Type ATPases, structural information is used from both ATP7A and ATP7B. These two can be used interchangeably to illustrate the overall structure of P(1B)-Type ATPases because they contain a sequence identity of greater than 80% [1]. Actuator Domain (A-Domain) and Membrane Domain (M-Domain)The for both ATP7A and ATP7B are 119 amino acid residues in length, and their actuator domain structure was solved using Nuclear Magnetic Resonance spectroscopy (PDB code 2kij) [1]. The structure is made up of which form two beta sheets [1]. The are packed together and form a distorted double-stranded beta-helix fold [1]. The two sheets are made up of ani-parallel beta strands 3, 4, 7 and 9 and 5, 6 and 8 respectively [1]. There are which flank the two beta sheets, and form a beta hairpin structure [1]. The final secondary structural element of the A-Domain are , which are extensions of the fourth and fifth transmembrane helices present in the M-Domain [1]. In summary, there are . The two alpha helices are extensions of the transmembrane helices of the overall protein, located at the N and C-terminus of the A-Domain respectively. The Actuator Domain of both ATP7A and ATP7B primarily serves a regulatory function [1]. The A-Domain contains the which is a threonine, glycine and glutamine residue [1]. This motif is necessary for the dephosphorylation of the P-Domain of the protein, therefore inactivating it [1]. Thus, the A-Domain's regulatory function is in the phosphatase step of the whole catalytic cycle of the ATPase [1]. The Membrane Domain of both ATP7A and ATP7B, which is also known as the transmembrane region of the proteins consists of contains eight transmembrane helices [1]. These transmembranous alpha helices main function is to provide an intramembranous channel for the Cu(I) to pass through the membrane [1]. The secondary function of the M-Domain is to anchor the whole ATPase into the plasma membrane, holding all of the other domains of the protein in proper orientation to one another [1]. The P-, and N-Domains of the Cu(I) transporting ATPase are located between the sixth and seventh transmembrane helices, and the A-Domain is located between the fourth and fifth [1]. Unfortunately, there is no individually solved structure for the P(1B)-Type ATPases ATP7A and ATP7B M-Domain, but in the solved structure for the A-Domain, an extension of the transmembrane helices from the M-Domain can be observed. The Membrane Domain of both ATP7A and ATP7B also contains a lengthy N-Terminus tail, which is approximately 70 amino acid residues in length [1]. This N-Terminal tail is structurally a part of the M-Domain of the proteins, but also contains its own unique function. The N-Terminal tail contains six copper binding sites in the Eukaryotic P(1B)-Type Cu(I) Transporting ATPases [1]. The primary function of this N-Terminal tail is to bind copper, and orient the copper for subsequent translocation through the intramembranous channel created by the eight transmembrane helices of the M-Domain [1]. A secondary function for the N-Terminus tail has been proposed as well; it is thought to be involved in a regulatory role through copper dependent interactions with other domains of the copper transporting proteins ATP7A and ATP7B [1]. Nucleotide-Binding Domain (N-Domain) and Phosphorylation Domain (P-Domain)The of the P(1B)-Type Copper(I) Transporting ATPases is approximately 17-kDa, whose primary function is to bind Adenine-Based nucleotides [5]. The structure was solved using Nuclear Magnetic Resonance spectroscopy, with and without bound ATP (PDB code 2arf) [5]. The N-Domain showed a marked increase in stability when ATP was bound, but exhibited no ATPase activity, concluding that the N-Domain is only the site for ATP binding [5]. The N-Domain consists of two types of secondary structure. The first is a , which consists of six anti-parallel beta strands [5]. The second type of secondary structural element are , which are found on either side of the central beta sheet [5]. In Summary, the N-Domain of the P(1B)-Type ATPases consists of a on either side of it. As stated above, the Nucleotide-Binding Domain's function is to bind ATP for subsequent phosphorylation of the P-Domain, which will enable the Cu(I) to be transported through the intramembranous channel formed by the M-Domain [5]. The N-Domain has a distinct coordination site for nucleotide binding which consists of residues [5]. These residues are in close proximity to each other, and are actually the most frequent residues mutated in a dysfunctional Cu(I) transporting ATPase, suggesting that this is in fact the location of ATP binding for the P(1B)-Type Cu(I) ATPases [5]. The Phosphorylation Domain, or P-Domain of ATP7A and ATP7B is located in between the sixth and seventh transmembrane helices of the M-Domain, in close proximity to the N-Domain of the proteins [1]. The primary function of the P-Domain is to accept the gamma phosphate that is removed from an N-Domain bound ATP [1]. The gamma phosphate binds to an Aspartate residue of the P-Domain located in the DKTG mofif [1]. Upon successful binding of the gamma phosphate, the P-Domain induces a conformational change in the transmembrane (M-Domain) region of the Cu(I) transporting protein, allowing successful release of the Cu(I) metal on to the other side of the membrane [1]. Catalytic Mechanism
Roles Within the CellCopper AbsorbtionCopper is mainly absorbed into the human body through the intestinal mucosa which is a process that is tightly regulated [6]. Once the copper has been absorbed into the body, it is transported to the liver through the blood stream bound to albumin and/or transcuperein [6]. Once copper in the blood stream has reached the liver, its uptake into hepatocytes is regulated by the protein CTR1 and the divalent metal transporter DMT1 [6]. After copper has been absorbed into the cell, the metallochaperone ATOX1 picks up copper from CTR1 [6]. At this point, depending on if total copper levels in the body are low or high, the copper will take one of two pathways; either delivery to ATP7B for distribution throughout the body and subsequent interaction with ATP7A, or delivery to ATP7B for biliary excretion. ATP7AThe P(1B)-Type Copper Transporting ATPase ATP7A is expressed in the majority of tissue in the body except for hepatocytes and its primary function is the delivery of copper to copper dependent proteins. Under low or normal copper levels within the cell ATP7A is located in the membrane of the trans-golgi network [6]. ATP7A's primary role is to receive copper from the metallochaperone ATOX1, and use the energy from ATP hydrolysis to transport the copper across the trans-golgi network's membrane into the tran-golgi network. This allows for coppers integration into copper-dependent enzymes such as lysyl oxidase [6]. Thus, under normal copper levels in the body ATP7A mediates and regulates copper transport into the trans-golgi network of non-hepatocyte cells and its subsequent integration into copper-dependent proteins. Under high copper levels in the body, ATP7A's is relocated within the cell, and its primary function changes. In response to high intracellular copper levels, ATP7A is relocated from the trans-golgi network to the plasma membrane of the cell [6]. ATP7A's relocation to the plasma membrane is an ATP dependent process, and has been shown to not require the synthesis of new proteins [6]. Once relocated to the plasma membrane, ATP7A proceeds to eliminate excess intracellular copper using its ATP dependent copper transport across the plasma membrane and out of the cell to ultimately eliminate high copper levels within the body. Because ATP7A is not expressed in the liver, its primary roll in copper homeostasis is to direct copper for integration into copper dependent proteins or for excretion from non-liver tissue. It is important to note this is only after copper has already been distributed throughout the body by ATP7B expressed in the liver. ATP7BThe P(1B)-Type Copper Transporting ATPase ATP7B is expressed primarily in hepatocytes of the liver. When the hepatocytes have low or normal intracellular copper levels, ATP7B is localized to the membrane of the trans-golgi network of the hepatocytes [6]. ATP7B receives intracellular copper from the metallochaperone ATOX1, and transports the copper into the trans-golgi network for its subsequent delivery to ceruloplasmin (for copper delivery throughout the body) or for biliary excretion [6]. Consequently, ATP7B's primary function is to transport copper into the trans-golgi network of hepatocytes only for its secretion to ceruloplasmin for bodily distribution or for biliary excretion from the body. ATP7B undergoes a relocation very similar to that of ATP7A when copper levels have reached an high level within heptatocytes. When copper concentration within hepatocytes is high, ATP7B is relocated to the plasma membrane of the hepatocytes [6]. ATP7B's relocation is done in the same manner as ATP7A; it is an ATP dependent process which does not require synthesis of new ATP7B proteins [6]. After ATP7B is relocated, its primary function is the transport of copper out of the hepatocytes for bodily excretion [6]. As a result of ATP7B only being expressed in hepatocytes, its primary roll is to supply the remainder of the bodies tissue with copper or discard copper through biliary excretion. In regards to ATP7B it is important to note that it does not distribute copper to copper dependent enzymes, rather it ultimately regulates the distribution of copper to ATP7B in non-liver tissue. SummaryCopper is absorbed into the body in the small intestines and then delivered to the liver through the blood stream. Once in the hepatocytes, copper is delivered to ATP7B for biliary excretion in the case of sufficient copper levels in the body, or for further distribution to non-liver tissues throughout the body. Copper is transported from the liver through the blood stream to bodily tissue where it absorbed by cells. Once absorbed into non-liver cells, copper is delivered to ATP7A for distribution to copper-dependent proteins, or for excretion from the body. MutationsCopper homeostasis within the body needs to be kept in tight regulation. This is because while copper is necessary for many biological functions within the body, free copper can have extremely detrimental consequences. Because ATP7A and ATP7B are both central proteins in copper homeostasis and regulation within the body, any mutation in either of these proteins which causes them to be dysfunction has severe consequences on health and copper homeostasis within the body. Menkes Disease and ATP7AMenkes disease is characterized by a general copper defficiency in the human body, and is a direct result of mutations present in ATP7A. Because ATP7A is responsible for the distribution of copper to many copper dependent enzymes within the body, a dysfunctional ATP7A results in subsequent malfunctioning of many copper dependent proteins [7]. Clinical features of Menkes Disease are commonly severe mental defects, growth retardation, laxity of skin and joints, hypopigmentation, and kinky or steely like hair [7]. All of these clinical features can be related back to a deficiency in copper. Unfortunately, in cases of Menkes Disease patients usually die within 3 years of life, but if diagnosed early there are some treatments available for the disease [7]. If diagnosed early in life, physicians need to take into account the following four consideratios: First, the block of copper absorption must be bypassed, second, treatment needs to be started as early as possible, third, circulating copper must be delivered to the brain and finally, copper needs to be made available within cells for copped dependent biosynthesis [7]. The only current treatment for Menkes Disease is copper replacement therapy in which copper-histidine (naturally occuring copper-amino acid) is administered to the patient [7]. The majority of mutations in the ATP7A gene that lead to Menkes disease are estimated to be de novo mutations [7]. Among these mutations are deletion/insertions, nonsense mutations, missense mutations, and splice site mutations which all occur at relatively equal frequency de novo to cause Menkes Disease [7]. Currently there is no cure for Menkes Disease. Wilsons Disease and ATP7BWilsons Disease is characterized by a copper overload in the human body, and is a result of mutations present in ATP7B. ATP7B is the protein primarily responsible for copper transport for biliary excretion and copper distribution throughout the body, and because of this, when ATP7B is dysfunctional there is no copper excretion from the body or distribution to throughout the body resulting in a copper overload in the liver. Clinical characterization of Wilsons disease includes toxic copper accumulation in the liver, brain and kidneys leading to hepatic abnormalities, progressive liver failure, neurological defects, and psychiatric symptoms [7]. Wilsons Disease has been known to present itself at many different ages, but does not lead to mortality at a young age like Menkes Disease. As a result, the diagnosis of Wilsons Disease varies in age and can usually be confirmed only after presentation of one or more of the symptoms above [7]. The treatment of Wilsons Disease focuses on two aspects: First, copper excretion from the body must be promoted especially in the liver and brain, and second, copper absorption from the diet must be regulated [7]. The main treatment for Wilsons Disease is with copper chelating agents such as penacillamine and trientine which will effectively remove copper from the body [7]. The majority of mutations in ATP7B gene which cause Wilsons Disease are missense mutations. In fact, missense mutations are responsible for over 60% of all Wilsons Disease cases [7]. Of these missense mutations the most prevalent are H1069Q in Europe and North America and R778L in Asia [7]. Currently there is no cure for Wilsons Disease, although patients who received liver transplants have experienced much less severe consequences of the disease. This is possibly because ATP7B is primarily expressed in the liver. | ||||||||||||
3D structures of ATPase
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 1.22 1.23 1.24 1.25 1.26 1.27 1.28 1.29 1.30 1.31 1.32 1.33 Banci L, Bertini I, Cantini F, Migliardi M, Natile G, Nushi F, Rosato A. Solution structures of the actuator domain of ATP7A and ATP7B, the Menkes and Wilson disease proteins. Biochemistry. 2009 Aug 25;48(33):7849-55. PMID:19645496 doi:10.1021/bi901003k
- ↑ 2.0 2.1 2.2 Dierick HA, Ambrosini L, Spencer J, Glover TW, Mercer JF. Molecular structure of the Menkes disease gene (ATP7A). Genomics. 1995 Aug 10;28(3):462-9. PMID:7490081 doi:http://dx.doi.org/10.1006/geno.1995.1175
- ↑ 3.0 3.1 3.2 Terada K, Schilsky ML, Miura N, Sugiyama T. ATP7B (WND) protein. Int J Biochem Cell Biol. 1998 Oct;30(10):1063-7. PMID:9785470
- ↑ 4.0 4.1 4.2 4.3 Kuhlbrandt W. Biology, structure and mechanism of P-type ATPases. Nat Rev Mol Cell Biol. 2004 Apr;5(4):282-95. PMID:15071553 doi:http://dx.doi.org/10.1038/nrm1354
- ↑ 5.0 5.1 5.2 5.3 5.4 5.5 5.6 5.7 Dmitriev O, Tsivkovskii R, Abildgaard F, Morgan CT, Markley JL, Lutsenko S. Solution structure of the N-domain of Wilson disease protein: distinct nucleotide-binding environment and effects of disease mutations. Proc Natl Acad Sci U S A. 2006 Apr 4;103(14):5302-7. Epub 2006 Mar 27. PMID:16567646
- ↑ 6.00 6.01 6.02 6.03 6.04 6.05 6.06 6.07 6.08 6.09 6.10 6.11 6.12 La Fontaine S, Mercer JF. Trafficking of the copper-ATPases, ATP7A and ATP7B: role in copper homeostasis. Arch Biochem Biophys. 2007 Jul 15;463(2):149-67. Epub 2007 May 7. PMID:17531189 doi:10.1016/j.abb.2007.04.021
- ↑ 7.00 7.01 7.02 7.03 7.04 7.05 7.06 7.07 7.08 7.09 7.10 7.11 7.12 de Bie P, Muller P, Wijmenga C, Klomp LW. Molecular pathogenesis of Wilson and Menkes disease: correlation of mutations with molecular defects and disease phenotypes. J Med Genet. 2007 Nov;44(11):673-88. Epub 2007 Aug 23. PMID:17717039 doi:10.1136/jmg.2007.052746
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, David Canner, Matthew Halstead, Alexander Berchansky